 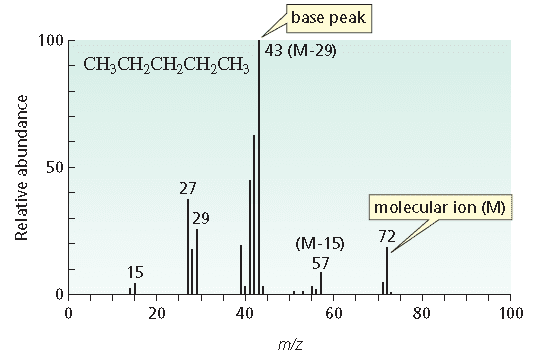
If there's any sulfurs or halogens in your molecule, they'll show up in the M+2 if there is one. The M+2 peak is similar to the M+1 in that it is 2 atomic mass units greater than the M+ peak. From the M+1 peak we can usually determine the number of carbons, oxygens, and nitrogens in the molecule using their known isotope abundances. The M+1 peak would just be the peak with an atomic mass 1 greater than the M+ peak. This is more or less your molecular weight of your molecule. It's essentially the entire molecule, minus a single electron, turning it into what we call a radical cation. The M+ peak is the largest atomic weight peak referred to as the molecular ion peak. If you're just sending a sample with a single element in it like Sal's example then you use that to find the isotopes and their abundances.īut if you're doing something in organic chemistry where we send organic compounds with multiple carbons of various chain lengths then we usually look at the M+, M+1, and M+2 peaks. I don't have experience using mass spectrometry so I don't actually know the process they used, but from my experience in AP Statistics that is the type of process we would use.ĭepends on which type of MS you're doing. If anything I said here is incorrect, please someone let me know. If K (our number of samples) is small then we wouldn't be as accurate as if we take more samples. Take multiple samples, use mass spectrometry to to find the relative abundances of each isotope, and then write that down.Ĭontinue this process for n=K samples, & then I can calculate the mean relative abundances for each isotope. Though I obviously did not find the relative abundances of the elements myself, here is what I have done. If we just take one sample & use the percentages from that one sample, then I would think that isn't the best inference for all of nature, though we can still make a claim for it. 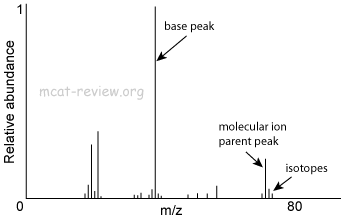
Whenever we take a random sample in nature & use mass spectrometry, we are finding the percentage that different isotopes occur in a sample of that element. In statistics, we can go around, take samples, and make an inference from it. We also analyzed statistics on adducts from spectra contained in the three selected mass spectral libraries.  
While the included adduct database focuses on adducts typically detected during liquid chromatography-mass spectrometry analyses, users may supply their own lists of adducts and charge states for calculating expected m/ z. This tool is intended to help researchers create identification libraries to collect evidence for the presence of molecules in mass spectrometry data. The calculator relies on user-selected subsets of the combined database to calculate expected m/ z for adducts of molecules supplied as formulas. MSAC includes a database of 147 potential adducts and adduct/neutral loss combinations and their mass-to-charge ratios ( m/ z) as extracted from the NIST/EPA/NIH Mass Spectral Library (NIST17), Global Natural Products Social Molecular Networking Public Spectral Libraries (GNPS), and MassBank of North America (MoNA). Here, adduct refers to a version of a parent molecule that is charged due to addition or loss of atoms and electrons resulting in a charged ion, for example, +. We describe the Mass Spectrometry Adduct Calculator (MSAC), an automated Python tool to calculate the adduct ion masses of a parent molecule. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
